Case 5: Tacticity Control (Atactic Polystyrene)
Case 5: Tacticity Control (Atactic Polystyrene)
Section titled “Case 5: Tacticity Control (Atactic Polystyrene)”In polymers with bulky side chains like polystyrene (PS), stereoregularity (tacticity) significantly affects packing and physical properties.
In fbtk, you can generate an atactic (random stereoregularity) polymer chain as an independent Molecule object using the Molecule.from_polymer method.
Target System
Section titled “Target System”- Polymer: Polystyrene (PS)
- Composition:
- Degree of Polymerization (DP): 20
- Count: 10 chains
- Tacticity: Atactic
- Total Atoms: 3,220
- Initial Target Density: 0.10 g/cm³ (Low-density start)
Calculation Code (run.py)
Section titled “Calculation Code (run.py)”This is the recommended configuration for creating an atactic chain with Molecule.from_polymer and starting relaxation from low density.
import fbtk
# 1. Create monomer from SMILESmonomer = fbtk.Molecule.from_smiles("*C(C*)c1ccccc1", name="PS")
# 2. Generate an Atactic polymer chain# Use from_polymer to create a single Molecule object with specified tacticitychain = fbtk.Molecule.from_polymer(monomer, degree=20, tacticity="atactic")
# 3. Initialize the Builder (Specify low density of 0.10)builder = fbtk.Builder(density=0.10)
# 4. Add the generated polymer chains to the systembuilder.add_molecule(chain, count=10)
# 5. Build the system and perform relaxationsystem = builder.build()print("Starting relaxation for Atactic Polystyrene...")system.relax()
# 6. Output resultschain.to_file("ps_atactic_chain.mol") # Individual atactic chain generatedsystem.to_file("ps_atactic_cell.mol2") # Relaxed unit cellprint("Done.")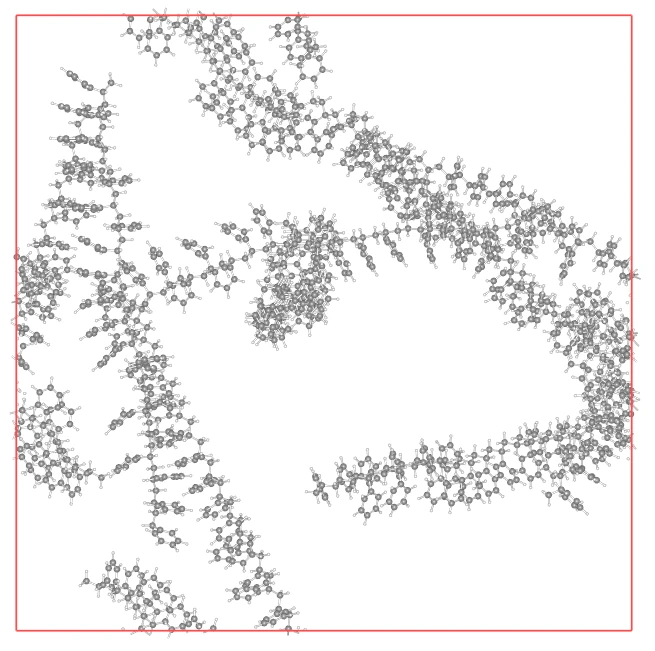
Reviewing the Execution Log
Section titled “Reviewing the Execution Log”The combination of a low-density start and atactic configuration allows even large-scale polystyrene systems to converge smoothly.
Atoms: 3220 | Bonds: 3410-------------------------------------------------------------------------------- | Fmax | FRMS | Total E |Iter | (kcal/mol/Å) | (kcal/mol/Å) | (kcal/mol) | Status-------------------------------------------------------------------------------- 0 | 42578.2095 | 6442.3968 | 1422137162651.3857 | 10 | 10670.1938 | 495.5003 | 570159.9917 | 20 | 5032.0412 | 224.4168 | 340325.8947 | ... 980 | 39.2758 | 0.9008 | 38346.5101 | 990 | 775.1007 | 16.9985 | 38246.3019 |--------------------------------------------------------------------------------=== Optimization Finished ===Reason: Max-IterTotal Time: 37.916s (Avg: 37.916ms / step)Final Energy: 38217.2511 kcal/molFinal FRMS: 1.5753 kcal/mol/ÅMin Distance: 1.0775 Å (Atoms 1473 and 1481)--------------------------------------------------------------------------------Result Explanation
Section titled “Result Explanation”Generation of Atactic Chains
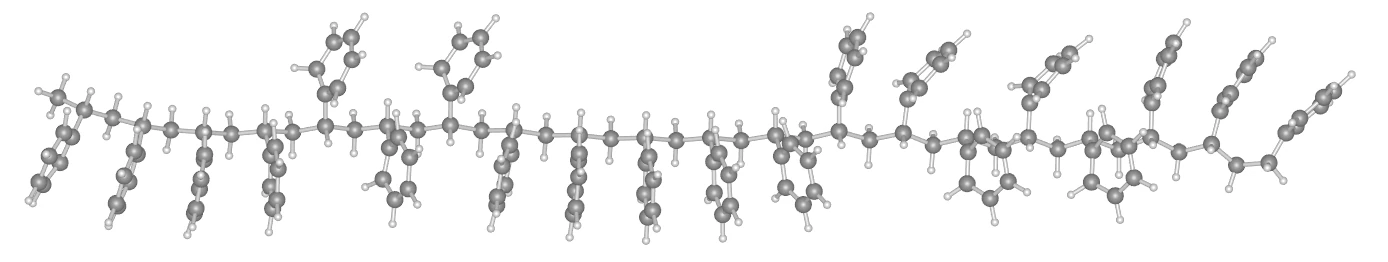
Section titled “Generation of Atactic Chains”Using Molecule.from_polymer constructs a concrete Molecule object in memory where side-chain orientations are randomly assigned to each asymmetric carbon in the backbone. By outputting this with to_file, you can also verify the 3D structure of a single chain before packing.
Improved Relaxation Quality
Section titled “Improved Relaxation Quality”Compared to structures where side chains are all aligned in the same direction (isotactic-like), atactic structures disperse steric hindrance more easily, allowing for smoother relaxation after packing. In this result, the Min Distance is 1.0775 Å, which is a good initial structure for a large-scale system.