Quickstart
Quickstart
Section titled “Quickstart”Basic Workflow of FBTK
Section titled “Basic Workflow of FBTK”Structure building in FBTK follows these three essential steps. Understanding this flow will make your building process smooth.
graph LR
A[Molecule<br/>Template Definition] --> B[Builder<br/>Packing & Placement]
B --> C[System<br/>Physics, Analysis, Export]
style A fill:#f9f,stroke:#333,stroke-width:2px
style B fill:#bbf,stroke:#333,stroke-width:2px
style C fill:#bfb,stroke:#333,stroke-width:2px
- Molecule: This is your “template” or “blueprint.” You define a single molecular structure from SMILES or files. Even a single polymer chain is generated as one
Moleculeobject here. - Builder: This is the “placer.” You specify “which molecules and how many” to pack into a box with a target density, and then execute the actual
build(). - System: This is the “realized system.” It contains the final coordinates and unit cell information (PBC). You can perform structural relaxation, RDF analysis, or save it to a file from here.
Quick Building from SMILES
Section titled “Quick Building from SMILES”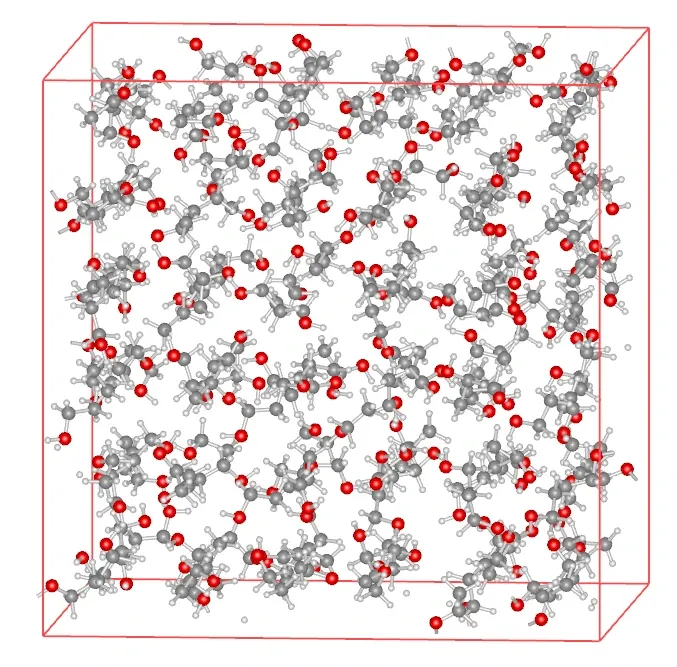
import fbtk
# 1. Define Molecule (Template)ethanol = fbtk.Molecule.from_smiles("CCO", name="Ethanol")
# 2. Placement with Builderbuilder = fbtk.Builder(density=0.789) # Specify target density (g/cm3)builder.add_molecule(ethanol, count=200) # Add 200 instances of the templatesystem = builder.build() # Execute placement
# 3. Operations on System (Realized)system.relax() # Structural relaxationsystem.to_file("initial.mol2") # Save results
# (Optional) Conversion to ASE Atoms# atoms = system.to_ase()# atoms.write("initial.data", format="lammps-data")